Computational Biochemistry Group
Head: Prof. Dr. Birgit Strodel
Forschungzentrum Jülich
IBI-7: Structural Biochemistry
Research
Amyloid aggregation: functional and disease related
 The aggregation of amyloid proteins can be abberant and leading to disease or functional and contributing to normal biological processes. The elucidation of the abberant aggregation mechanisms is a key step in preventing the onset of many degenerative diseases such as Alzheimer's or Parkinson's disease. Moreover, understanding the reason for the lack of toxicity of functional amyloids is essential for the design of new biocompatible nanomaterials. Using atomistic computer simulations we explore the aggregation of peptides and proteins with focus on the structure of small order oligomers known to exhibit toxic activity. To understand the kinetics of aggregation we use transition networks in combination with first passage time distributions and free energy disconnectivity graphs.
The aggregation of amyloid proteins can be abberant and leading to disease or functional and contributing to normal biological processes. The elucidation of the abberant aggregation mechanisms is a key step in preventing the onset of many degenerative diseases such as Alzheimer's or Parkinson's disease. Moreover, understanding the reason for the lack of toxicity of functional amyloids is essential for the design of new biocompatible nanomaterials. Using atomistic computer simulations we explore the aggregation of peptides and proteins with focus on the structure of small order oligomers known to exhibit toxic activity. To understand the kinetics of aggregation we use transition networks in combination with first passage time distributions and free energy disconnectivity graphs.
Contact: Bogdan Barz,
Martin Carballo Pacheco,
Birgit Strodel
Recent publications:
M. Carballo-Pacheco, A. Ismail, and B. Strodel. Oligomer Formation of Toxic and Functional Amyloid Peptides Studied with Atomistic Simulations J. Phys. Chem. B, DOI: 10.1021/acs.jpcb.5b04822 (2015)
[link]
B. Barz, O. Olubiyi, and B. Strodel.
Early amyloid β-protein aggregation precedes conformational change
Chem. Commun., 50, 5373-5375 (2014)
[link]
B. Barz, D.J. Wales, and B. Strodel.
A Kinetic Approach to the Sequence–Aggregation Relationship in Disease-related Protein Assembly
J. Phys. Chem. B., 118, 1003-1011 (2014)
[link]
Amyloid-lipid interactions
 The etiology of Alzheimer's disease is thought to be linked to interactions between the amyloid β-peptide (Aβ) and neural cell membranes. Membrane disruption and increased ion conductance have been observed in vitro in the presence of Aβ, and it is assumed that the same phenomena occur in the brain of Alzheimer's patients. The effects of Aβ on lipid behavior have been characterized experimentally, but structural and causal details are lacking. Simulations of Aβ or other amyloid proteins in a bilayer environment may provide the information necessary to explain their toxicity. We follow different simulation approaches in order to gain a better understanding of the interactions between amyloid proteins and lipid bilayers.
The etiology of Alzheimer's disease is thought to be linked to interactions between the amyloid β-peptide (Aβ) and neural cell membranes. Membrane disruption and increased ion conductance have been observed in vitro in the presence of Aβ, and it is assumed that the same phenomena occur in the brain of Alzheimer's patients. The effects of Aβ on lipid behavior have been characterized experimentally, but structural and causal details are lacking. Simulations of Aβ or other amyloid proteins in a bilayer environment may provide the information necessary to explain their toxicity. We follow different simulation approaches in order to gain a better understanding of the interactions between amyloid proteins and lipid bilayers.
Contact: Michael Owen,
Birgit Strodel
Recent publications:
C. Poojari, D. Xiao, V. S. Batista and B. Strodel.
Membrane permeation induced by aggregates of human islet amyloid polypeptides
Biophys. J., 105, 2323-2332 (2013)
[link]
C. Poojari, A. Kukol and B. Strodel.
How the amyloid-β peptide and membranes affect each other: an extensive simulation study.
BBA-Biomembranes, 1828, 327-339 (2013)
[link]
Effects of Cu(II) ions and oxidation on amyloid-β aggregation
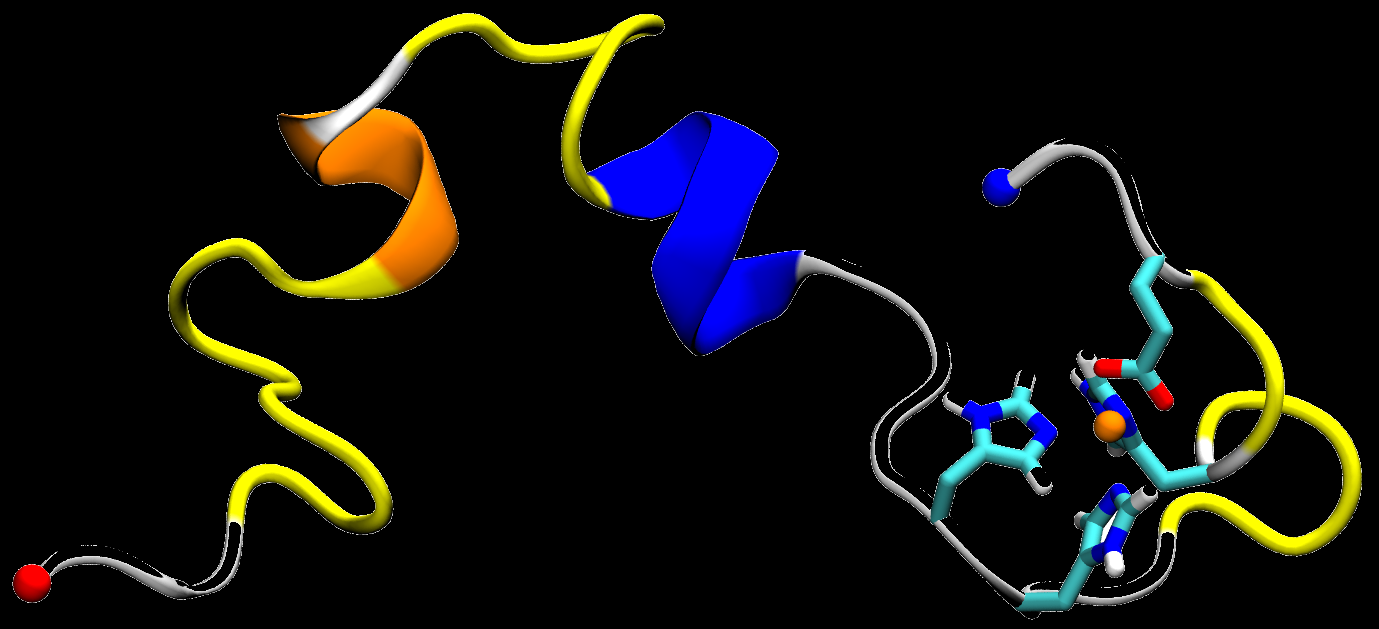 Metal ions such as Cu2+ have been proposed to play a key role in the formation of amyloid-β plaques. Cu2+ can bind to
Aβ directly and modulate its aggregation process. As a redox reactive metal ion, copper is also crucial for the production of reactive oxygen species (ROS)
and oxidative stress, which further contributes to the pathology of Alzheimer's disease.
While the coordination and interactions between copper and Aβ were elucidated experimentally and computationally, little information is
available about the conformational dynamics of the copper-bound Aβ oligomers. In our group, QM/MM calculations and molecular dynamics simulations are applied
to determine the effect of Cu-binding and ROS production on the aggregation of Aβ, and on the subsequent ROS-initiated oxidation of lipids and proteins central to the pathology of Alzheimer's disease.
Metal ions such as Cu2+ have been proposed to play a key role in the formation of amyloid-β plaques. Cu2+ can bind to
Aβ directly and modulate its aggregation process. As a redox reactive metal ion, copper is also crucial for the production of reactive oxygen species (ROS)
and oxidative stress, which further contributes to the pathology of Alzheimer's disease.
While the coordination and interactions between copper and Aβ were elucidated experimentally and computationally, little information is
available about the conformational dynamics of the copper-bound Aβ oligomers. In our group, QM/MM calculations and molecular dynamics simulations are applied
to determine the effect of Cu-binding and ROS production on the aggregation of Aβ, and on the subsequent ROS-initiated oxidation of lipids and proteins central to the pathology of Alzheimer's disease.
Contact:
Qinqhua Liao,
Michael Owen,
Birgit Strodel
Recent publications:
Q. Liao, S.C.L. Kamerlin, and B. Strodel.
Development and Application of a Nonbonded Cu2+ Model That Includes the Jahn–Teller Effect
J. Phys. Chem. Lett., 6, 2657–2662 (2015)
[link]
Protein dynamics: linking theory and experiment
 The aim of this project is to establish close links between MD simulations and experimental observables
as obtained from NMR, FRET or FCS speactroscopic measurements. To this end we closely collaborate
with the groups of Dieter Willbold and Philipp Neudecker (NMR, Heinrich Heine University Düsseldorf and
Forschungszentrum Jülich) and Claus Seidel (FRET/FCS, Heinrich Heine University Düsseldorf).
The two proteins we have studied thus far are the autophagy-related proteins GATE-16 and GABARAP.
The aim of this project is to establish close links between MD simulations and experimental observables
as obtained from NMR, FRET or FCS speactroscopic measurements. To this end we closely collaborate
with the groups of Dieter Willbold and Philipp Neudecker (NMR, Heinrich Heine University Düsseldorf and
Forschungszentrum Jülich) and Claus Seidel (FRET/FCS, Heinrich Heine University Düsseldorf).
The two proteins we have studied thus far are the autophagy-related proteins GATE-16 and GABARAP.
Contact:
Oliver Schillinger,
Birgit Strodel
Recent publications:
Ma, P. et al.
Interaction of Bcl-2 with the Autophagy-Related Protein GABARAP -- Biophysical Characterization and Functional Implications
J. Biol. Chem., 113, 1-26 (2013)
[link]
Ma, P.; Schillinger, O. et al..
Conformational Polymorphism in Autophagy-Related Protein GATE-16.
Biochemistry, DOI: 10.1021/acs.biochem.5b00366 (2015)
[link]
Computational enzyme design
 Enzymes are prone to changes in the tertiary structure due to the oxidative and thermal stress and other chemical reactions. The catalytic activity and selectivity of an enzyme are hugely affected by its 3D structure. Even modest structural changes usually lead to severe catalytic activity loss, making an enzyme inconvenient for industrial applications. On the other side, certain structural changes, purposely applied to the enzyme, can increase its oxidative and thermal stability and catalytic activity. Larger changes can even alter the specificity to a substrate so the enzyme can be forced to catalyze a reaction on a non-natural, industrially important substrate. In collaboration with experimental groups, we use a multiscale approach for enzyme rational design: from all-atom MD simulations to hybrid QM/MM calculations.
Enzymes are prone to changes in the tertiary structure due to the oxidative and thermal stress and other chemical reactions. The catalytic activity and selectivity of an enzyme are hugely affected by its 3D structure. Even modest structural changes usually lead to severe catalytic activity loss, making an enzyme inconvenient for industrial applications. On the other side, certain structural changes, purposely applied to the enzyme, can increase its oxidative and thermal stability and catalytic activity. Larger changes can even alter the specificity to a substrate so the enzyme can be forced to catalyze a reaction on a non-natural, industrially important substrate. In collaboration with experimental groups, we use a multiscale approach for enzyme rational design: from all-atom MD simulations to hybrid QM/MM calculations.
Contact:
Dusan Petrovic,
Birgit Strodel
Structure prediction of proteins and protein complexes
 The determination of protein structures is one of the most important challenges in biochemistry. Computational techniques help to find the three-dimensional arrangement
of atoms. However, the exact determination of the experimental structure from a denaturated or unfolded protein is still a challenge. The use of structural restraints obtained from experiments like NMR measurements show significant improvement in this field of research.We combined of the basin-hopping approach to global optimization with NMR chemical shift restraints.
We currently work on the extension of the basin-hopping approach towards a protein-protein docking approach.
Here, we employ the OPEP coarse-grained force field, which in comparison to all-atom force fields
smooths the potential energy surface and increase calculation speed.
To allow for flexibility during docking we use local Monte Carlo moves at the binding site.
The determination of protein structures is one of the most important challenges in biochemistry. Computational techniques help to find the three-dimensional arrangement
of atoms. However, the exact determination of the experimental structure from a denaturated or unfolded protein is still a challenge. The use of structural restraints obtained from experiments like NMR measurements show significant improvement in this field of research.We combined of the basin-hopping approach to global optimization with NMR chemical shift restraints.
We currently work on the extension of the basin-hopping approach towards a protein-protein docking approach.
Here, we employ the OPEP coarse-grained force field, which in comparison to all-atom force fields
smooths the potential energy surface and increase calculation speed.
To allow for flexibility during docking we use local Monte Carlo moves at the binding site.
Contact:
Philipp Kynast,
Birgit Strodel
Recent publications:
F. Hoffmann, I. Vancea, S.G. Kamat, and B. Strodel.
Protein structure prediction: assembly of secondary structure elements by basin-hopping
Chem. Phys. Chem., 15, 3378–3390 (2014)
[link]
F. Hoffmann and B. Strodel.
Protein structure prediction using global optimization by basin-hopping with NMR shift restraints.
J. Chem. Phys., 138:025102 (2013)
[link]
Menu
News
Amyloid-β peptide dimers undergo a random coil to β-sheet transition in the aqueous phase but not at the neuronal membrane
H. Fatafta, M. Khaled, M. C. Owen, A. Sayyed-Ahmad, B. Strodel
Proc. Nat. Acad. Sci. USA 118, e2106210118 (2021)
Energy landscapes of protein aggregation and conformation switching in intrinsically disordered proteins
B. Strodel
J. Mol. Biol. 433, 167182 (2021)